
CRC1238 Data Publication Guidelines
The CRC is committed to the principles of reproducibility, scientific integrity, and open access.
For the publication of our results, we will adhere to the following guidelines (last update: Jan. 2024).
- To make all publications of the CRC available and freely accessible worldwide, we are using arXiv. All the information required to reproduce scientific results is included in the publication and the accompanying (data-)supplements.
- Simultaneously with the paper, we publish the underlying data either directly with the journal or via Zenodo. We publish at least the data underlying all figures (incl. appendices) in accessible formats (minimal standard). Best practice is to publish also all relevant raw data and, if applicable, scripts, configuration files and codes and all information needed to process the raw data. Codes shared on GitHub can also be linked to Zenodo.
Examples of published data can be accessed via the CRC 1238 community on Zenodo.
Guideline for publishing data with Zenodo
To publish research data we recommend to do the following before submitting a paper to arXiv or a journal.
The following text is partially based on the ML4Q publishing guide and Refs. [1,2,3].
1. Data preparation
1.1
Check whether the journal requires or provides the option to publish the electronic research data with the journal. If yes, follow the guidelines of the journal, if not, proceed with preparing your data for Zenodo. Even in the first case, you may find the following text useful.
1.2
Prepare a folder with all relevant data on your computer. This folder will later be uploaded to the repository. Organize the folder in a logical and understandable manner, avoiding deep folder hierarchies. In most cases you can use the structure of the paper to organize the data, e.g., by creating a subfolder for each figure (incl. appendices).
Create a readme.txt file in the root folder that describes the relevant data/codes/configuration files, where to find them, and (if applicable) what needs to be done to reproduce the results.
1.3
Include at least the data which has directly been used in the publication. This means, that every figure in your publication should be accompanied by the extracted data in a format that is readable to others. For example, if there is a color plot, the underlying data array should be published. If possible, aim to export your data in a common format, for example csv or hdf5 files. Avoid data formats that need proprietary software to view. It is acceptable to upload the data in another format, as long as that data file is accompanied by an instruction on how to load the data.
1.4
On top of the data described above, discuss with your coauthors what other data is useful to share. Best practice is to publish all raw data, all custom-made codes and all relevant scripts and configuration files of instruments and codes together with a description of how the data is processed (e.g., in the readme file). Record the software packages that you used, including their versions. Include source codes and/or scripts you used to process the data. The goal is that others can reproduce the published results using the published codes and the measured raw data.
1.5
For the publication of code and scripts make it portable and usable by others. For example, do not read data with absolute paths (e.g., C:/my_name/PhD/project/raw_data/measurement.hd5), but only with relative paths (e.g., raw_data/measurement.hd5).
1.6
Double-check everything. Make sure that all coauthors and other relevant persons (e.g., authors of codes you want to publish) have agreed to the publication of the data, scripts, and codes. Remove all unnecessary files, non-shareable data objects (raw and processed!), passwords hardcoded in your scripts, comments containing private information, and so on.
1.7
Finally, make a single archive file from your data folder. It is recommended to use zip, as it is supported by practically every operating system natively. Your data is now ready to be published.
2. Upload to Zenodo
2.1
If you do not yet have an ORCID identifier (a unique identifier for researchers required by many journals and universities), we recommend that you get one from orcid.org. Then create an account on zenodo.org. If you log in with your ORCID account, the ORCID is automatically linked to your account.
2.2
In the upper right corner, you find a button marked by + to create a new upload. You will get a page where you can upload your files, determine the type of upload, and create metadata for the research object. Fill out the basic information. For resource type, you may choose e.g. dataset. Choose a meaningful title, for example Data and code for “title of your publication”. Make sure to add all authors and affiliations. The description should contain information about what the data is and where to find which parts of the data. You can use the content of your readme file. Choose the correct license, the default Creative Commons Attribution 4.0 International is a good choice.
2.3
Save the draft, preview it and check everything carefully. After publication, the metadata can still be changed but not the submitted files, which are stored permanently. However, one can always upload a new version of the same project using the New version button.
2.4
Now you can publish the files. After publishing, you get a permanent DOI to the published data. If you want to change the data later, you have to upload a new version of the same project on Zenodo.
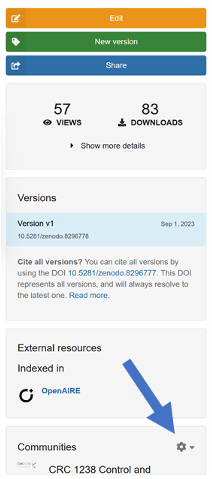
2.5
Finally, add the published data to the CRC 1238 community. Below the edit button (see Fig.), search for Communities, click on the wheel (arrow in the figure), search for the “CRC 1238” community and add your paper to the community.
3. Update preprints and publications
Your preprint and later the paper should cite the data repository. For example, Phys. Rev. recommends to add a sentence before the acknowledgements. You can use something like “The supporting data and codes for this article are available from Zenodo [REF].” Here [Ref] is an entry in your bibliography citing the DOI of the form AUTHOR NAMES, YEAR, Zenodo, https://doi.org/10.5281/zenodo.xxxxxx. All preprints and papers published within CRC 1238 should also be added to our publication database which can be found under crc1238.uni-koeln.de/publikationen/.
References:
[1] Akhmerov, A., & Steele, G. (2019). Open Data Policy of the Quantum Nanoscience Department, TU Delft. Zenodo. https://doi.org/10.5281/zenodo.2556949
[2 Kesteren, E.-J. (2017). A short practical guide for preparing and sharing your analysis code. https://odissei-data.nl/en/2022/06/a-short-practical-guide-for-preparing-and-sharing-your-analysis-code/
[3] D. Grothe (2024), ML4Q Publishing Guide, https://ml4q.de/rdm/publishing-guide/